There’s some big, positive news on the stem cell front today.
Two new innovative papers both by teams led by Sheng Ding of Gladstone Institutes with UCSF report all-chemical direct reprogramming of human somatic cells. Ding’s team took skin cells and by exposing them to cocktails of small molecules was able to turn them directly into precursors for heart muscle and neural cells. The two direct reprogramming papers were published in Science here and in Cell Stem Cell here. The former paper is Cao, et al. and the latter is Zhang, et al. These reports together are a very big deal.

It’s been almost a decade since Shinya Yamanaka first reported the creation of mouse induced pluripotent stem cells (IPSC) using a cocktail of four factors and only one year later, he and others reported the creation of human IPSC.
Almost all the subsequent reports of creation of IPSC have involved at least transient introduction of genes or proteins. Then in 2013 a team led by Hongkui Deng reported all-chemical production of mouse IPSC.
There’s also been great interest in so-called direct reprogramming (transdifferentiation), which is in a sense “skipping the middle man” of IPSC to turn somatic cells such as skin cells directly into another cell type such as neural or heart cells. However, direct reprogramming hasn’t been done chemically…until now with the work led by Ding.
You can see a field of mature neurons made from the chemically-induced neural precursors (ciNSC) above. Imagine in the future, therapies might be made from a patient’s own cells that are directly reprogrammed into the desired neural, heart, or other types of cells.
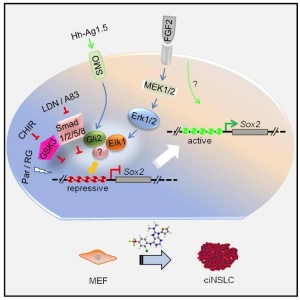
I’ve included the graphical abstract showing the process for neural induction.
This neural direct reprogramming was accomplished with so-called M9 medium containing 9 chemicals consisting overall of:
50% Neural basal, 50% DMEM/F12/Glutamax, 1x N2, 1x B27 without vitamin A, 0.075% BSA, 0.1 mM nonessential amino acids, with 3 mM CHIR99021, 100 nM LDN193189, 0.5 mM A83-01, 0.5 mM Hh-Ag1.5, 1 mM retinoic acid, 10 mM SMER28, 10 mM RG108, 2 mM Parnate, and 10 ng/ml bFGF.
The recipe for the chemical induction media for cardiac cells, which is called 9C, also contains 9 compounds:
CHIR99021 (GSK3 inhibitor, activate Wnt pathway), A83-01 (TGFbeta receptor inhibitor, inhibits TGFbeta signaling), BIX01294 (histone methyltransferase inhibitor, modulates epigenetics), AS8351 (histone demethylase inhibitor, also modulates epigenetics), SC1 (ERK inhibitor), Y27632 (ROCK inhibitor), OAC2 (target unknown, modulates epigenetics), SU16F (PDGF receptor inhibitor) and JNJ10198409 (PDGF receptor inhibitor).
Figure 5f from this paper is shown below at least outlining the process of 9C induction. The neural induction was done in mouse cells, while the cardiac was done in human cells.
I need to take a deeper read of these two papers, but my initial impression is that they will have a big, positive impact.
Some open questions do remain. As best as I could tell, for instance, there is no discussion of mutations in the chemically-induced cells, tumorigenesis. Whole genome sequencing data will be needed. I would also like to see more data on DNA methylation. I expect we will learn more on these areas in follow up studies.
A press release from The Gladstone quoted Ding about his big picture view of the papers’ impact:
“This method brings us closer to being able to generate new cells at the site of injury in patients,” said Ding, the senior author on both studies. “Our hope is to one day treat diseases like heart failure or Parkinson’s disease with drugs that help the heart and brain regenerate damaged areas from their own existing tissue cells. This process is much closer to the natural regeneration that happens in animals like newts and salamanders, which has long fascinated us.”
As Ding alludes too, with chemical reprogramming in theory a future treatment might stimulate a human patient’s own endogenous cells to change type in vivo. I wonder if to some extent could that already be happening in us at times based on our bodies’ own chemicals? Does an aberrant form of this contribute to cancer? Hal Weintraub’s pioneering work on MyoD promoting muscle cell differentiation of other cell types comes to mind too.
Overall, these new direct reprogramming papers are a big step in the right direction to bring direct reprogramming forward and into the translational pipeline.
Does this mean that cells has to continuously maintain their chromatin to remain committed to their lineage, or does the cocktail mimic some fetal development related signalling?
I have a feeling that we know so little about histone modification status of different cell lines, for example was there ever published a concise investigation on whole DNA histone modification? I believe without such studies we are just playing lottery.
Its the chromatin , stupid!
Dr. Knoepfler, would you consider a longer or indefinite open comment period for your posts? My comment is more germane to another post (http://j.mp/bioheart-2015) but comments are closed for that one.
Anyway, writing last year in ‘Bioheart on the edge in 2015’ you highlighted a lawsuit. That suit has been settled according to the company’s annual report filed March 28th (http://j.mp/us-stem-ar-2015). As an investor I regard the terms of the settlement favorably. It’s just one of many factors that goes into evaluating what we can hope for in the future though.
I was little surprised that you punctuated the post back then with that point. I find your scientific skepticism very welcome in the discourse around the regulatory “sweet spot.” But the now settled lawsuit had little to do with science or regulation.
I am not a scientist, my interest stems particularly from having diabetic peripheral neuropathy. Does this development mean that sometime in the next few years I might get my feet back? I really want to get my feet back.
does anyone outside of sheng ding’s lab actually use (or publish on) these protocols? My sense was no, but I could be mistaken…
Is any indication given that the components of 9C could be used in vivo? Perhaps to steer regenerating cardiac stem cells toward cardiomyocyte lineage instead of scar tissue?
From skin to heart/muscle seems like a big change. I wondered if bone to/from cartilage might be easier. (Excuse my preoccupation with cartilage.) Then I realized that there is a great big catch to all of this. Changing a cell is only a tiny part of remodelling/fixing tissue. Here is a really neat link where you can find a user-controlled animation that takes you through all the steps involved in bone remodelling:
https://depts.washington.edu/bonebio/ASBMRed/growth.html
On one level it’s sort of humbling. On another level it’s a great big reminder about just how messy evolution is compared to intelligently designed machines.
This brings me to a whimsical, philosophical point. Scientists tend to be design-oriented with respect to how we do things… (OK, maybe more whimsical than philosophical. Philosophers would do better.)
Paul,
The work was done with mouse cells, or human?
Neural is mouse, cardiac human.